In today’s society, when we are curious about our family lineage, we can upload our DNA to genetic testing platforms to provide insight into our ancestral origins. These platforms can estimate where an individual’s ancestors may have lived and can identify genetic connections to populations around the world. However, an important question remains: how far back can these methods realistically trace human ancestry, and what limitations prevent us from fully connecting modern populations to early humans?
A common method of learning about ancestral ties is by analyzing variants in an individual’s genetic code and where they occur most commonly in the world. This is done by comparing a large subset of an individual’s DNA code variants (most commonly single-nucleotide variants (SNVs)) frequencies to then be matched to a geographic region in the world (Jorde and Bamshad 2020). The identification of the geographic location with the highest frequency of a certain variant defines the area to be the most likely location of an ancestor with said variant. To assess these variants, methods involving assays using DNA microarray and DNA sequencing is used to analyze these variants. When the SNV is found, it is then matched to the location with the highest frequency of this variant. As seen in Figure 1, the overall percentage of said variants in the sample DNA can then determine the percentage of an individual’s ancestry from different regions across the world (Jorde and Bamshad 2020).
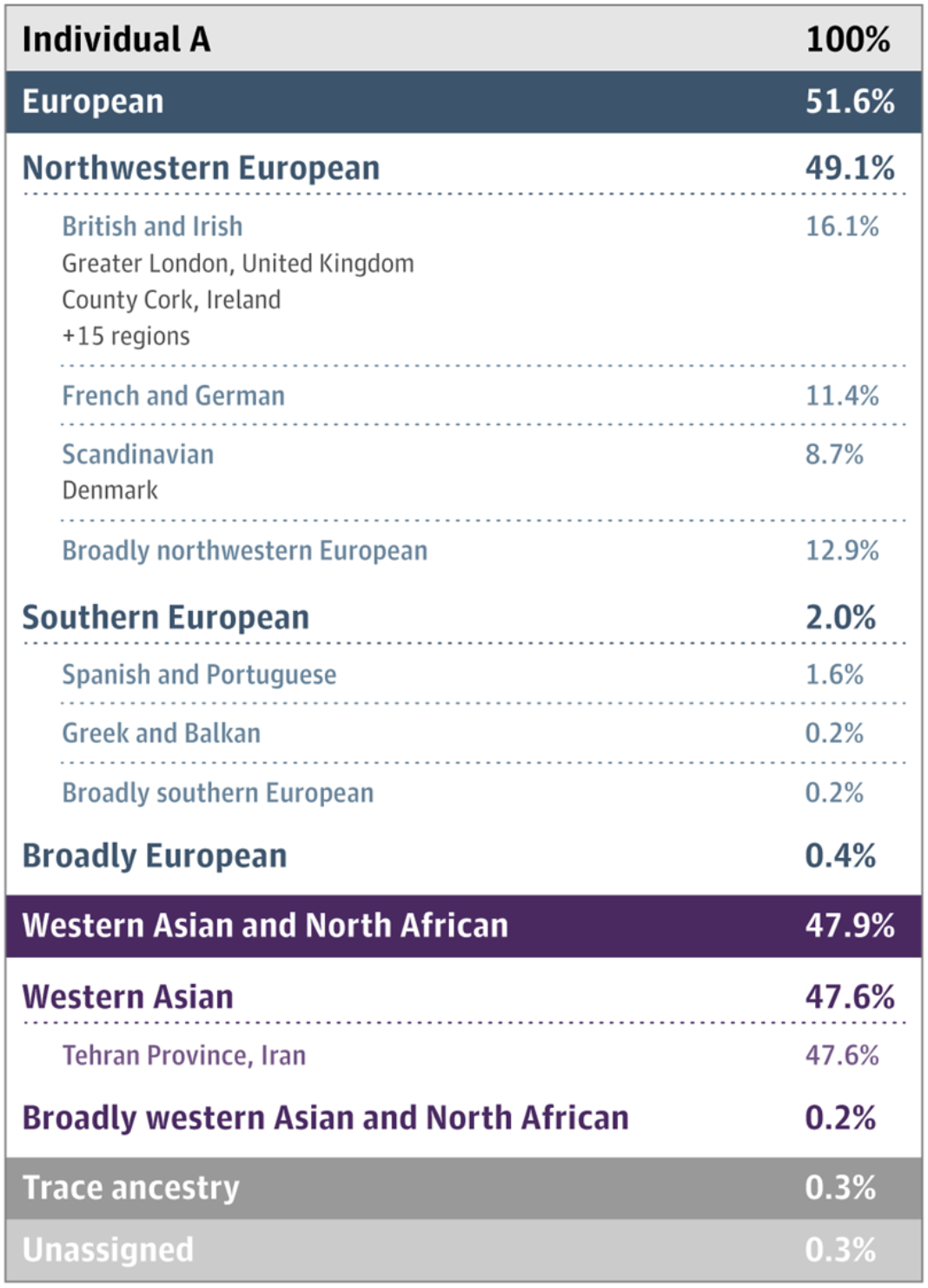
Figure 1. Hypothetical ancestral results of an individual who sampled their DNA to be studied. The different percentages of geographic origin can be seen, and through the results, the highest percentage is found in Northwestern and southern Europe, whereas the rest of the identifiable region is Western Asia and North Africa (Jorde and Bamshad 2020).
While relatively accurate to finish local family portraits, ancestral tracking through DNA has several limitations when looking at ancestral heritage to early humans. Certain problematic areas consist of time duration, record accuracy, as well as regional and socioeconomic bias. When attempting to connect an individual back to early human populations, it can be highly inaccurate as genealogical records decay with each generation. This makes DNA insufficient for studying deep ancestral roots (Bird et al 2025). In addition to their decaying accuracy through time, they are not neutral sources as they are reflections of individual understandings, which are vulnerable to bias. The information is also input by individuals, meaning there is large room for error due to misspellings, incorrect data, and incomplete documentation (Hatton 2021). Along with these flaws that impact results, the differences of socioeconomic status can also be a factor in the recognition of ancestry in certain regions of the world, leaving many communities unrepresented (Nelson 2008).
Overall, the reconstruction of ancestral linkage is used across the world to provide answers to those with questions about lineage and genetic diseases. These methods allow for the reconnections between lost relatives and provide the opportunity to learn more about family history and culture. However, when attempting to connect today’s population with early humans, we are greeted with several challenges, time being the main factor. Since records decay with time, an accurate depiction of a family tree throughout the entire human timeline is highly unlikely. These flaws, however, have the potential to lead to newer methods in the future, so maybe we will all have a framed portrait of an extensive family tree hanging on our walls one day.
References
Bird, Nancy et al. “Power and Limitations of Inferring Genetic Ancestry.” Annals of human genetics vol. 89,5 (2025): 264-273. doi:10.1111/ahg.70007
Hatton, Stephen B. “Genealogy’S Assumptions about Written Records and Originality.” Genealogy, vol. 5, no. 1, 2021, https://doi.org/10.3390/genealogy5010021.
Jorde, Lynn B, and Michael J Bamshad. “Genetic Ancestry Testing: What Is It and Why Is It Important?” JAMA vol. 323,11 (2020): 1089-1090. doi:10.1001/jama.2020.0517
Nelson, Alondra. “Bio science: genetic genealogy testing and the pursuit of African ancestry.” Social studies of science vol. 38,5 (2008): 759-83. doi:10.1177/0306312708091929
Leave a Reply
You must be logged in to post a comment.